What are Protein Domains?
Protein domains often correspond to structural domains which are self-stabilizing and fold independently of the rest of the protein chain. They may occur independently or as part of a complex multi-domain protein [1]. Protein domains can therefore function independently from the rest of the protein chain. Domains ultimately determine overall protein function.
Protein motifs are super-secondary structures such as alpha helices or beta sheets that don't necessarily have a biological function like domains. They are not independent structures. Motifs are short segments found within domains that collectively create/build them [2].
Protein motifs are super-secondary structures such as alpha helices or beta sheets that don't necessarily have a biological function like domains. They are not independent structures. Motifs are short segments found within domains that collectively create/build them [2].
Domains of SYNGAP1
There are multiple programs that can be utilized to characterize proteins based on their sequences and identify which domains and motifs are present within the protein. It is important to realize different programs create different protein domain structures. For SYNGAP1, Pfam and SMART display different domains. Pfam does not recognize the PH (Pleckstrin homology) domain whereas SMART does. SMART displays miniature unidentified regions while Pfam demonstrates one large segment (DUF3498) and lacks a coiled coil region.
Biological Functions of SYNGAP1 Domains
Pleckstrin Homology Domain |
The PH domains are small, modular and appear in a wide variety of proteins. The domain family possesses multiple functions including the abilities to bind inositol phosphates, and various proteins. Through these interactions, PH domains play a role in recruiting proteins to different membranes, thus targeting them to appropriate cellular compartments or enabling them to interact with other components of the signal transduction pathways [3].
C2 Domain
The C2 domain is involved in targeting proteins to cell membranes. Often times the C2 domains contain a Ca+2 binding motif, but not all. They appear to bind phospholipids, inositol polyphosphates, and intracellular proteins. Within the SYNGAP1 protein the C2 domain is seen to play a role in GAP activity which is a unique property not commonly seen for this domain [4].
RasGAP Domain
Ras proteins are membrane-associated molecular switches that bind GTP and GDP and slowly hydrolyze GTP to GDP. This intrinsic GTPase activity of Ras is regulated by a family of proteins collectively known as 'GAP' or GTPase-activating proteins. The RasGAP domain of SYNGAP1 is a GTPase-activator protein for Ras-like GTPases. It's an all alpha-helical domain that accelerates the GTPase activity of Ras, thereby "switching" it into an "off" position [3].
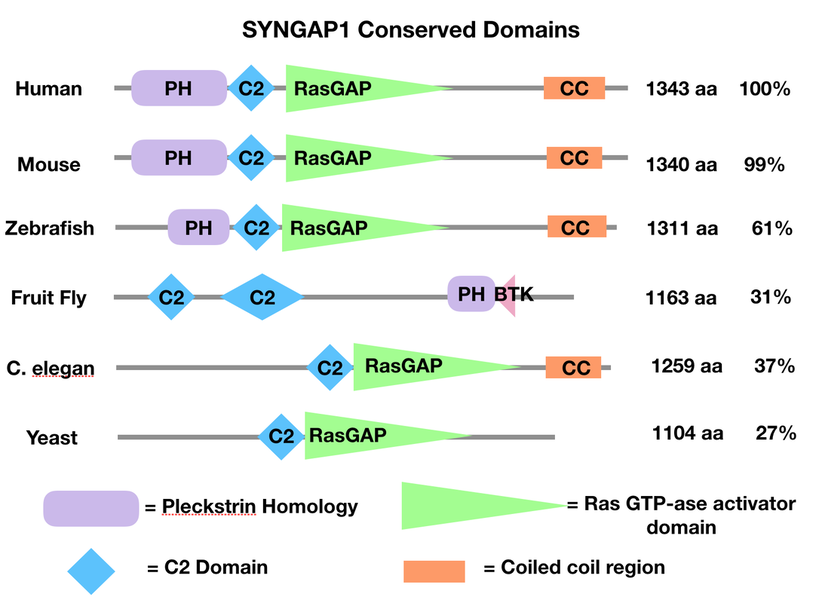
Which Domains Are Conserved in SYNGAP1?
Discussion
After understanding the conserved domains of SYNGAP1 across species specifically our model organism, Danio rerio (zebrafish) we can manipulate these domains and observe the different resulting phenotypes.
References
[1] UniProt Domain Overview. Retrieved from https://www.uniprot.org/help/domain
[2] EMBL-EBI Training Center. Retrieved from https://www.ebi.ac.uk/training/online/course/biomacromolecular-structures-introduction-ebi-reso/proteins/structural-motifs
[3] http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=128104153129266241552686486thabMMWMTF
[4] Pena, Vladimir et al. “The C2 domain of SynGAP is essential for stimulation of the Rap GTPase reaction” EMBO reports vol. 9,4 (2008): 350-5. Retrieved from https://www.researchgate.net/publication/5528195_The_C2_domain_of_SynGAP_is_essential_for_stimulation_of_the_Rap_GTPase_reaction
Figure 1: http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=7233267301751552613810ixHeodtjYx
Figure 2: http://pfam.xfam.org/search/sequence
SYNGAP1 Conserved Domains created by Abigail Jaquish
[2] EMBL-EBI Training Center. Retrieved from https://www.ebi.ac.uk/training/online/course/biomacromolecular-structures-introduction-ebi-reso/proteins/structural-motifs
[3] http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=128104153129266241552686486thabMMWMTF
[4] Pena, Vladimir et al. “The C2 domain of SynGAP is essential for stimulation of the Rap GTPase reaction” EMBO reports vol. 9,4 (2008): 350-5. Retrieved from https://www.researchgate.net/publication/5528195_The_C2_domain_of_SynGAP_is_essential_for_stimulation_of_the_Rap_GTPase_reaction
Figure 1: http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=7233267301751552613810ixHeodtjYx
Figure 2: http://pfam.xfam.org/search/sequence
SYNGAP1 Conserved Domains created by Abigail Jaquish
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone at University of Wisconsin- Madison.