Protein interaction networks are diagrams depicting all proteins that interact within a biochemical event and/or electrostatic forces that serve a distinct biological function as a complex. The protein interactome describes the full repertoire of a biological system’s protein–protein interactions. Proteins are often nodes connected to other proteins by lines between them creating a web like structure. Networks can be designed for a single protein or for the entire network of a biological process. Creating protein interaction networks is a method to better understand and predict a proteins wide range of roles and functions [1].
Interaction Network for SYNGAP1
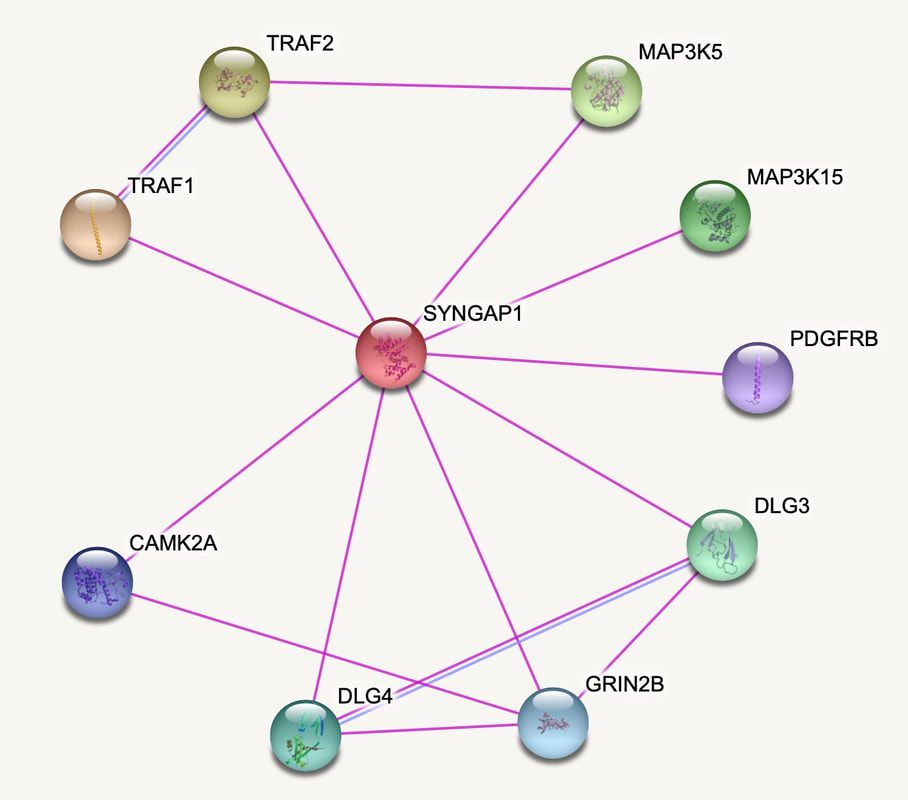
Interaction networks can be built using various programs such as STRING, IntAct or BioGrid. To create SYNGAP1's network I utilized the STRING database seen below. The network was built based on data from previous experiments.
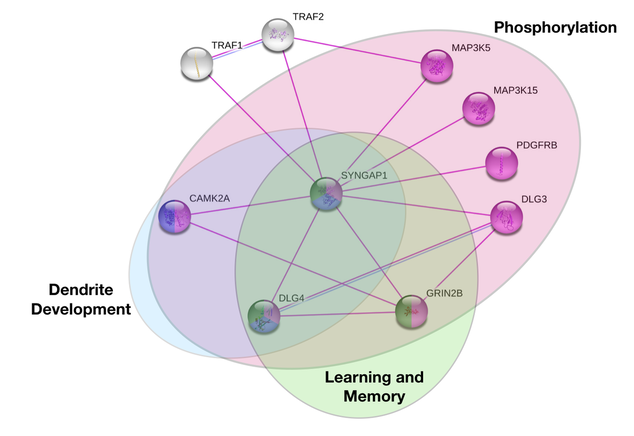
After further analysis within STRING or in other databases including GeneOntology or Panther a more informational depiction of the protein interactions can be created by separating proteins based on their ontology. SYNGAP1 has many functions within neurons, but when deciphering which proteins particularly play a role in behavior the network decreases.
Discussion
It is clear from SYNGAP1's interaction networks and highlighted regions of biological interest that it plays a role in many different processes. The proteins within the network are also a part of many more biological functions outside of the ones highlighted. Ultimately demonstrating that SYNGAP1 plays a vital role in the regulation of many processes and it is a difficult protein to study as it effects a wide range of aspects.
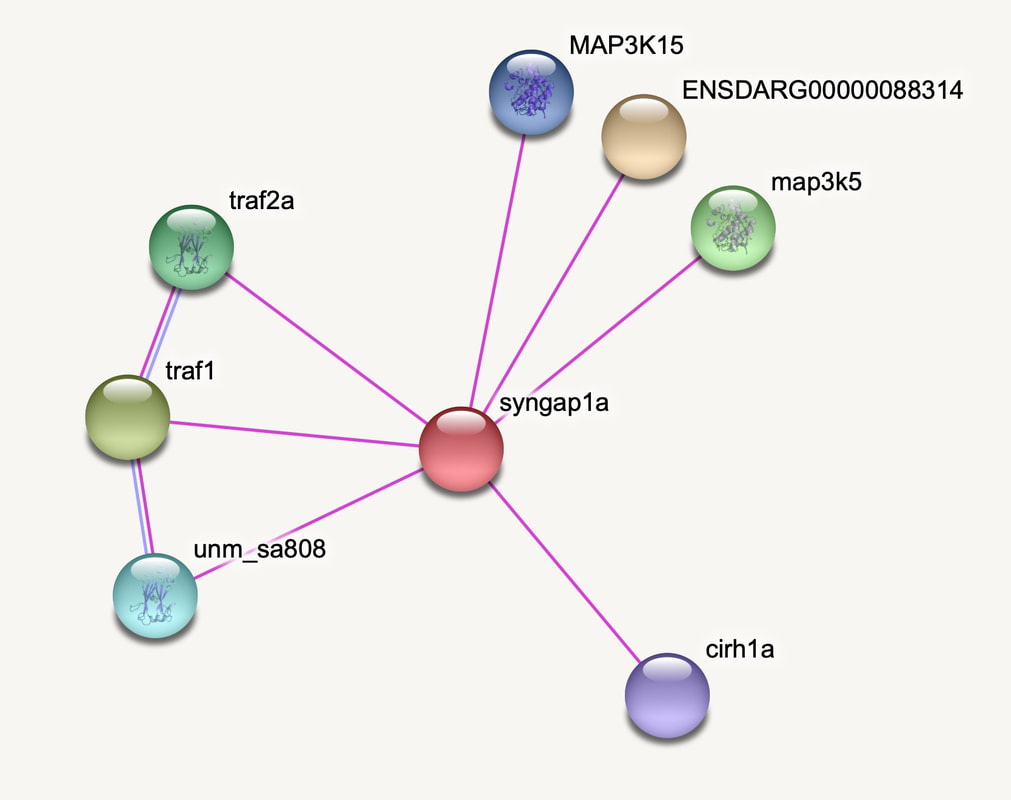
The interaction network for zebrafish SYNGAP1 was much more difficult to apply gene ontology too because the proteins are much less characterized and aren't listed to possess biological functions. Our study will help develop the biological role of the proteins that interact with SYNGAP1 in a zebrafish model.
The interaction network for zebrafish SYNGAP1 was much more difficult to apply gene ontology too because the proteins are much less characterized and aren't listed to possess biological functions. Our study will help develop the biological role of the proteins that interact with SYNGAP1 in a zebrafish model.
References
[1] https://www.nature.com/subjects/protein-protein-interaction-networks
Images:
Header: https://ese224.seas.upenn.edu/connections/
Figure 1: https://string-db.org/cgi/network.pl?taskId=DkjmhLy6bhcs
Figure 2: https://string-db.org/cgi/network.pl?taskId=q0RXaIaboPrM
Figure 3: Made by Abby Jaquish
Images:
Header: https://ese224.seas.upenn.edu/connections/
Figure 1: https://string-db.org/cgi/network.pl?taskId=DkjmhLy6bhcs
Figure 2: https://string-db.org/cgi/network.pl?taskId=q0RXaIaboPrM
Figure 3: Made by Abby Jaquish
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone at University of Wisconsin- Madison.